Code
library(causatr)
library(tinyplot)
data("nhefs")
nhefs_complete <- nhefs[!is.na(nhefs$wt82_71) & !is.na(nhefs$education), ]static() — set treatment to a fixed value
shift() — add a constant to observed treatment
scale_by() — multiply observed treatment by a factor
threshold() — clamp treatment within bounds (g-comp only)dynamic() — deterministic treatment rule
ipsi() — incremental propensity score intervention (IPW only)
stochastic() — user-defined randomised intervention (g-comp only)
NULL — the natural coursecausatr separates model fitting (causat()) from causal contrasts (contrast()). The intervention — the hypothetical change in treatment — is specified at contrast time, not fit time. This design lets you fit once and contrast under many different interventions without re-estimating nuisance models.
This vignette tours every intervention constructor, shows which estimators support it, and demonstrates the calling pattern on real and simulated data.
| Constructor | What it does | G-comp | IPW | AIPW | Matching |
|---|---|---|---|---|---|
static(v) |
Set treatment to v for everyone |
All types | Binary, categorical | All types | Binary |
shift(d) |
Add d to observed treatment |
Continuous | Continuous, count (integer d) |
Continuous | — |
scale_by(c) |
Multiply observed treatment by c |
Continuous | Continuous, count (integer-preserving) | Continuous | — |
threshold(lo, hi) |
Clamp treatment to [lo, hi] |
Continuous | — | — | — |
dynamic(rule) |
Apply a user-defined deterministic rule | All types | Binary, categorical | All types | — |
ipsi(d) |
Multiply odds of treatment by d |
— | Binary | — | — |
stochastic(f) |
Draw treatment from user-supplied distribution | All types | — | — | — |
grace_period(w) |
Delay treatment initiation by w periods (natural-history MTP) |
Longitudinal, discrete | — | — | — |
carry_forward() |
Carry the baseline treatment forward (LOCF; natural-history MTP) | Longitudinal, discrete | — | — | — |
NULL |
Natural course (observed values) | All types | All types | All types | — |
grace_period() and carry_forward() are natural-history modified treatment policies (G-LMTPs): the intervened treatment at time \(t\) depends on the counterfactual natural-value history of treatment, so they cannot be written as dynamic() rules (which see only the observed lag). They are estimated by an augmented-data sequential regression and require a longitudinal g-computation fit with a discrete treatment — see the longitudinal vignette for runnable examples.
All examples use the NHEFS dataset (Hernán & Robins, 2025):
qsmk — quit smoking (0/1). Used for static(), dynamic(), and ipsi() examples.smokeintensity — cigarettes smoked per day. Used for shift(), scale_by(), and threshold() examples.wt82_71 — weight change in kg.static() — set treatment to a fixed valueThe simplest and most common intervention. static(1) means “everyone is treated”; static(0) means “no one is treated”. For categorical treatments, pass the level name as a string (e.g. static("B")).
fit_gc <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "qsmk",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeintensity + I(smokeintensity^2) + smokeyrs + I(smokeyrs^2) +
factor(exercise) + factor(active) + wt71 + I(wt71^2),
model_fn = stats::glm
)
contrast(
fit_gc,
interventions = list(quit = static(1), continue = static(0)),
reference = "continue",
type = "difference"
)Estimator: gcomp · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| quit | 5.21 | 0.421 | 4.384 | 6.035 |
| continue | 1.747 | 0.217 | 1.321 | 2.173 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| quit vs continue | 3.462 | 0.468 | 2.546 | 4.379 |
fit_ipw <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "qsmk",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeintensity + I(smokeintensity^2) + smokeyrs + I(smokeyrs^2) +
factor(exercise) + factor(active) + wt71 + I(wt71^2),
estimator = "ipw",
model_fn = stats::glm,
propensity_model_fn = stats::glm
)
contrast(
fit_ipw,
interventions = list(quit = static(1), continue = static(0)),
reference = "continue",
type = "difference"
)Estimator: ipw · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| quit | 5.288 | 0.447 | 4.412 | 6.163 |
| continue | 1.788 | 0.217 | 1.363 | 2.214 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| quit vs continue | 3.499 | 0.49 | 2.538 | 4.46 |
fit_aipw <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "qsmk",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeintensity + I(smokeintensity^2) + smokeyrs + I(smokeyrs^2) +
factor(exercise) + factor(active) + wt71 + I(wt71^2),
estimator = "aipw",
model_fn = stats::glm,
propensity_model_fn = stats::glm
)
contrast(
fit_aipw,
interventions = list(quit = static(1), continue = static(0)),
reference = "continue",
type = "difference"
)Estimator: aipw · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| quit | 5.254 | 0.44 | 4.391 | 6.116 |
| continue | 1.777 | 0.218 | 1.349 | 2.205 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| quit vs continue | 3.477 | 0.483 | 2.531 | 4.423 |
fit_match <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "qsmk",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeintensity + I(smokeintensity^2) + smokeyrs + I(smokeyrs^2) +
factor(exercise) + factor(active) + wt71 + I(wt71^2),
estimator = "matching",
estimand = "ATT"
)
contrast(
fit_match,
interventions = list(quit = static(1), continue = static(0)),
reference = "continue",
type = "difference"
)Estimator: matching · Estimand: ATT · Contrast: difference · CI method: sandwich · N: 403
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| quit | 4.525 | 0.435 | 3.672 | 5.378 |
| continue | 1.184 | 0.383 | 0.433 | 1.935 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| quit vs continue | 3.341 | 0.559 | 2.246 | 4.436 |
All four methods should give broadly similar estimates (methodological triangulation). Discrepancies signal model misspecification in one or more approaches.
shift() — add a constant to observed treatmentA modified treatment policy (MTP) that shifts each individual’s observed treatment by a fixed amount. For continuous treatments, this answers “what would happen if everyone’s exposure increased (or decreased) by delta units?”
fit_gc_cont <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "smokeintensity",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeyrs + I(smokeyrs^2) + factor(exercise) + factor(active) +
wt71 + I(wt71^2),
model_fn = stats::glm
)
res_shift_gc <- contrast(
fit_gc_cont,
interventions = list(reduce10 = shift(-10), observed = NULL),
reference = "observed",
type = "difference"
)
res_shift_gcEstimator: gcomp · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| reduce10 | 2.684 | 0.256 | 2.183 | 3.186 |
| observed | 2.638 | 0.199 | 2.248 | 3.028 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| reduce10 vs observed | 0.046 | 0.181 | -0.308 | 0.4 |
Under IPW, shift(delta) uses a density-ratio weight: the ratio of the treatment density evaluated at the inverse-shifted value to the density at the observed value. The pushforward sign matters — shift(delta) evaluates at A - delta, not A + delta — because the weight uses the inverse map.
fit_ipw_cont <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "smokeintensity",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeyrs + I(smokeyrs^2) + factor(exercise) + factor(active) +
wt71 + I(wt71^2),
estimator = "ipw",
model_fn = stats::glm,
propensity_model_fn = stats::glm
)
res_shift_ipw <- contrast(
fit_ipw_cont,
interventions = list(reduce10 = shift(-10), observed = NULL),
reference = "observed",
type = "difference"
)
res_shift_ipwEstimator: ipw · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| reduce10 | 2.69 | 0.248 | 2.205 | 3.175 |
| observed | 2.638 | 0.199 | 2.248 | 3.028 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| reduce10 vs observed | 0.051 | 0.163 | -0.268 | 0.371 |
est <- data.frame(
method = c("G-comp", "IPW"),
estimate = c(
res_shift_gc$contrasts$estimate[1],
res_shift_ipw$contrasts$estimate[1]
),
ci_lower = c(
res_shift_gc$contrasts$ci_lower[1],
res_shift_ipw$contrasts$ci_lower[1]
),
ci_upper = c(
res_shift_gc$contrasts$ci_upper[1],
res_shift_ipw$contrasts$ci_upper[1]
)
)
tinyplot(
estimate ~ method,
data = est,
type = "pointrange",
ymin = est$ci_lower,
ymax = est$ci_upper,
xlab = "Method",
ylab = "Difference in weight change (kg)",
main = "shift(-10): reduce smoking by 10 cigs/day"
)
abline(h = 0, lty = 2, col = "grey40")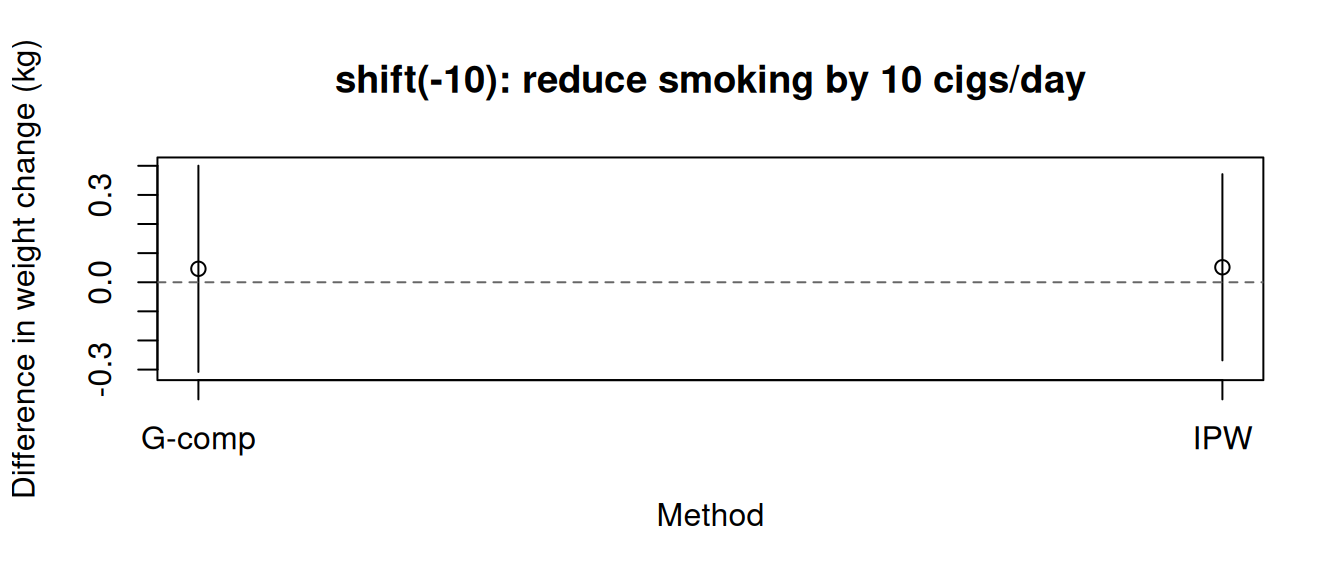
threshold() is not supported under IPW because the pushforward of a continuous density under a boundary clamp is a mixed measure (point masses at the bounds plus continuous density in the interior). Use estimator = "gcomp" for threshold interventions.
For integer-valued count treatments, pass propensity_family = "poisson" or "negbin" to use a count density model instead of the default Gaussian.
DGM. A Poisson-distributed count treatment (e.g. number of clinic visits) confounded by \(L\), with a linear outcome model.
\[ \begin{aligned} L &\sim N(0,1) \\ A &\sim \text{Poisson}\!\bigl(\exp(0.5\,L)\bigr) \\ Y &= 2 + 1.5\,A + L + \varepsilon, \quad \varepsilon \sim N(0,1) \end{aligned} \]
True contrast: \(\mathbb{E}[Y(A+1)] - \mathbb{E}[Y(A)] = 1.5\).
set.seed(6)
n <- 3000
L <- rnorm(n)
A <- rpois(n, exp(0.5 * L))
Y <- 2 + 1.5 * A + L + rnorm(n)
count_df <- data.frame(Y = Y, A = A, L = L)
fit_pois <- causat(
count_df,
outcome = "Y",
treatment = "A",
confounders = ~ L,
estimator = "ipw",
propensity_family = "poisson",
model_fn = stats::glm,
propensity_model_fn = stats::glm
)
res_pois <- contrast(
fit_pois,
interventions = list(plus1 = shift(1), observed = NULL),
reference = "observed",
type = "difference"
)
res_poisEstimator: ipw · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 3000
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| plus1 | 5.267 | 0.069 | 5.133 | 5.402 |
| observed | 3.692 | 0.049 | 3.595 | 3.788 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| plus1 vs observed | 1.576 | 0.05 | 1.478 | 1.674 |
The shift delta must be integer — shift(0.5) on a count treatment aborts because dpois() returns 0 at non-integer arguments.
scale_by() — multiply observed treatment by a factorA proportional MTP. scale_by(0.5) halves each individual’s exposure; scale_by(2) doubles it.
Estimator: gcomp · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| halved | 2.686 | 0.259 | 2.178 | 3.193 |
| observed | 2.638 | 0.199 | 2.248 | 3.028 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| halved vs observed | 0.047 | 0.185 | -0.316 | 0.411 |
Under IPW, scale_by(c) evaluates the density at A / c and includes the Jacobian |1/c| in the density-ratio weight.
Estimator: ipw · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| halved | 2.45 | 0.291 | 1.88 | 3.021 |
| observed | 2.638 | 0.199 | 2.248 | 3.028 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| halved vs observed | -0.188 | 0.223 | -0.625 | 0.249 |
threshold() — clamp treatment within bounds (g-comp only)Caps treatment at a ceiling and/or floor. threshold(0, 20) means “nobody smokes more than 20 cigarettes per day; those below 0 are set to 0”.
Estimator: gcomp · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| cap20 | 2.659 | 0.208 | 2.252 | 3.066 |
| observed | 2.638 | 0.199 | 2.248 | 3.028 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| cap20 vs observed | 0.021 | 0.08 | -0.137 | 0.178 |
This intervention is only available under g-computation. Under IPW, the density-ratio weight is not well-defined at the boundary clamp points (point-mass discontinuity in the pushforward density). causatr rejects threshold() under estimator = "ipw" with an informative error pointing at g-comp.
dynamic() — deterministic treatment ruleA dynamic intervention assigns treatment based on individual characteristics. The rule function receives two arguments — the full data and the observed treatment vector — and returns a vector of counterfactual treatment values.
Assign quitting only to heavy smokers (> 20 cigarettes/day at baseline):
Estimator: gcomp · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| rule | 2.725 | 0.204 | 2.325 | 3.124 |
| all_quit | 5.21 | 0.421 | 4.384 | 6.035 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| rule vs all_quit | -2.485 | 0.337 | -3.146 | -1.824 |
Under IPW, dynamic() on a binary treatment uses Horwitz–Thompson indicator weights: w_i = I(A_i = rule_i) / f(A_i | L_i). This is well-defined because the treatment takes only two values and each individual’s counterfactual is deterministic.
Estimator: ipw · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| rule | 2.803 | 0.355 | 2.107 | 3.499 |
| all_quit | 5.288 | 0.447 | 4.412 | 6.163 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| rule vs all_quit | -2.485 | 0.385 | -3.239 | -1.73 |
Dynamic rules on continuous treatments work under g-comp (predict-then-average on the outcome model). Here, heavy smokers are reduced to 20 while others keep their observed level:
Estimator: gcomp · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| capped | 2.659 | 0.208 | 2.252 | 3.066 |
| observed | 2.638 | 0.199 | 2.248 | 3.028 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| capped vs observed | 0.021 | 0.08 | -0.137 | 0.178 |
dynamic() on continuous treatment is not supported under IPW. A deterministic rule on a continuous treatment is a Dirac per individual, which has no Lebesgue density — the density ratio is not defined. Rewrite the rule as a smooth shift() or scale_by(), or use estimator = "gcomp".
ipsi() — incremental propensity score intervention (IPW only)Kennedy (2019)’s incremental propensity score intervention multiplies each individual’s odds of treatment by delta. It is a stochastic MTP that never materialises a specific counterfactual treatment value — instead, it shifts the entire propensity score distribution.
The closed-form weight is: \[w_i = \frac{\delta \cdot A_i + (1 - A_i)}{\delta \cdot p_i + (1 - p_i)}\]
where \(p_i = P(A = 1 \mid L_i)\) is the fitted propensity.
IPSI is binary-treatment, IPW-only — g-comp and matching cannot evaluate it because there is no counterfactual treatment column to predict at.
Estimator: ipw · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| double_odds | 3.125 | 0.218 | 2.697 | 3.552 |
| natural | 2.638 | 0.199 | 2.248 | 3.028 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| double_odds vs natural | 0.486 | 0.066 | 0.357 | 0.615 |
Estimator: ipw · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| half_odds | 2.284 | 0.197 | 1.898 | 2.669 |
| natural | 2.638 | 0.199 | 2.248 | 3.028 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| half_odds vs natural | -0.355 | 0.051 | -0.455 | -0.255 |
Varying delta traces out a dose-response curve on the odds scale:
deltas <- c(0.25, 0.5, 1, 2, 4)
ipsi_results <- lapply(deltas, function(d) {
res <- contrast(
fit_ipw,
interventions = list(shifted = ipsi(d), natural = NULL),
reference = "natural",
type = "difference"
)
data.frame(
delta = d,
estimate = res$contrasts$estimate[1],
ci_lower = res$contrasts$ci_lower[1],
ci_upper = res$contrasts$ci_upper[1]
)
})
ipsi_df <- do.call(rbind, ipsi_results)
tinyplot(
estimate ~ delta,
data = ipsi_df,
type = "pointrange",
ymin = ipsi_df$ci_lower,
ymax = ipsi_df$ci_upper,
xlab = expression(delta ~ "(odds multiplier)"),
ylab = "E[Y(IPSI)] - E[Y(natural)] (kg)",
main = "IPSI dose-response: effect on weight change"
)
abline(h = 0, lty = 2, col = "grey40")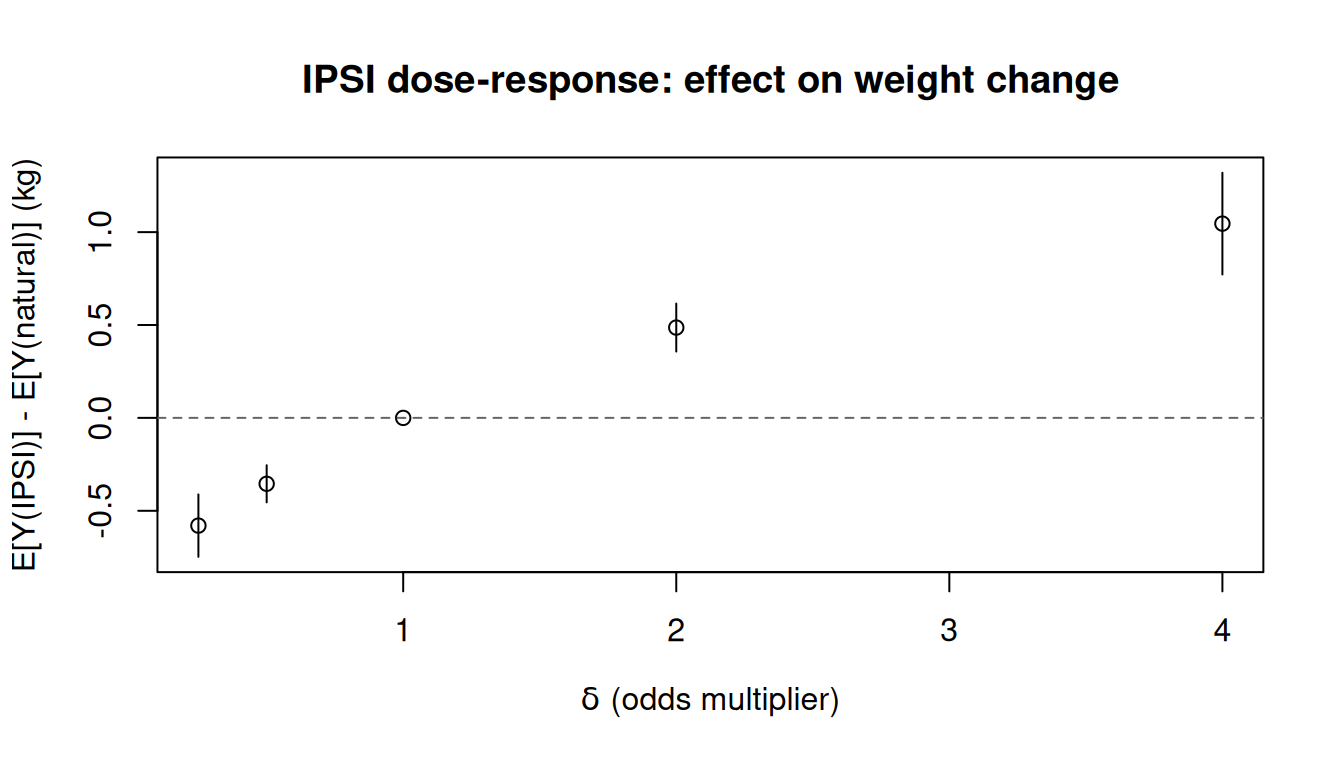
At delta = 1 the intervention is the natural course, so the contrast is zero. Larger delta makes quitting more likely and the effect on weight gain grows.
stochastic() — user-defined randomised intervention (g-comp only)A stochastic intervention draws each individual’s counterfactual treatment from a user-supplied distribution that may depend on covariates. Unlike dynamic(), which assigns a single deterministic value, stochastic() integrates over the randomness by Monte Carlo: the g-formula is evaluated n_mc times, each with an independent draw from the sampler, and the predictions are averaged.
The sampler function has the same signature as a dynamic() rule — function(data, treatment) — but should return an independent random draw each time it is called.
Assign quitting with probability that depends on smoking intensity:
Estimator: gcomp · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| random_quit | 3.489 | 0.242 | 3.014 | 3.963 |
| all_quit | 5.21 | 0.421 | 4.384 | 6.035 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| random_quit vs all_quit | -1.721 | 0.232 | -2.176 | -1.266 |
The estimate is a population average over both individuals and the stochastic policy.
Add a random perturbation (rather than a fixed shift) to each individual’s smoking intensity:
Estimator: gcomp · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| noisy_reduce | 2.661 | 0.211 | 2.248 | 3.074 |
| observed | 2.638 | 0.199 | 2.248 | 3.028 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| noisy_reduce vs observed | 0.023 | 0.09 | -0.154 | 0.2 |
Choosing n_mc: 100–500 is typical. The sandwich SE accounts for the MC draws used at estimation time, so there is no need to inflate n_mc just for inference. If point estimates visibly change when you re-run with a different seed, increase n_mc.
stochastic() is only supported under estimator = "gcomp" (point and longitudinal). IPW and matching cannot evaluate it because the density ratio of a user-defined stochastic policy is generally intractable.
NULL — the natural courseNULL means “do not intervene; use the observed treatment values”. It is the natural reference for MTPs:
Estimator: gcomp · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| reduce10 | 2.684 | 0.256 | 2.183 | 3.186 |
| halved | 2.686 | 0.259 | 2.178 | 3.193 |
| cap20 | 2.659 | 0.208 | 2.252 | 3.066 |
| observed | 2.638 | 0.199 | 2.248 | 3.028 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| reduce10 vs observed | 0.046 | 0.181 | -0.308 | 0.4 |
| halved vs observed | 0.047 | 0.185 | -0.316 | 0.411 |
| cap20 vs observed | 0.021 | 0.08 | -0.137 | 0.178 |
Both g-comp and IPW support multiple interventions in a single contrast() call. All pairwise differences against the reference are computed, and the variance-covariance matrix covers all interventions jointly.
Estimator: ipw · Estimand: ATE · Contrast: difference · CI method: sandwich · N: 1566
| intervention | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| reduce10 | 2.69 | 0.248 | 2.205 | 3.175 |
| halved | 2.45 | 0.291 | 1.88 | 3.021 |
| observed | 2.638 | 0.199 | 2.248 | 3.028 |
| comparison | estimate | se | ci_lower | ci_upper |
|---|---|---|---|---|
| reduce10 vs observed | 0.051 | 0.163 | -0.268 | 0.371 |
| halved vs observed | -0.188 | 0.223 | -0.625 | 0.249 |
The table below summarises which combinations are supported and which are rejected with an informative error. The restrictions reflect genuine architectural limits of the density-ratio framework, not implementation gaps.
| Intervention | G-comp (point) | G-comp (longitudinal) | IPW | AIPW | Matching |
|---|---|---|---|---|---|
static() |
All treatment types | All treatment types | Binary, categorical | All types | Binary only |
shift() |
Continuous | Continuous | Continuous, count (integer delta) | Continuous | — |
scale_by() |
Continuous | Continuous | Continuous, count (integer-preserving) | Continuous | — |
threshold() |
Continuous | Continuous | — (mixed pushforward) | — | — |
dynamic() |
All treatment types | All treatment types | Binary, categorical | All types | — |
ipsi() |
— | — | Binary only | — | — |
stochastic() |
All treatment types | All treatment types | — (intractable density ratio) | — | — |
NULL |
All | All | All | All | — |
Why some combinations are rejected:
threshold() under IPW: The pushforward of a continuous density under a boundary clamp is a mixed measure — point masses at lo/hi plus continuous density in the interior. The density ratio is not well-defined.dynamic() on continuous treatment under IPW: A deterministic rule on a continuous treatment is a Dirac per individual with no Lebesgue density.static(v) on continuous treatment under IPW: Nobody is observed exactly at v, so the HT indicator I(A = v) is zero almost surely.ipsi() under g-comp/matching: IPSI shifts the propensity, not the treatment value. There is no counterfactual treatment column to predict at.stochastic() under IPW/matching: The density ratio of an arbitrary user-defined stochastic policy is generally intractable. Only g-computation (Monte Carlo integration) supports it.static() / dynamic() / threshold() / ipsi() on count treatments: Count treatments (enabled via propensity_family = "poisson" or "negbin") use dpois() / dnbinom() for the density ratio. static() is degenerate (HT indicator is zero almost surely on a count distribution). dynamic() has the same Dirac problem as on continuous. threshold() has the same mixed- measure problem. ipsi() is Bernoulli-specific. Only shift() (integer delta) and scale_by() (where A / factor is integer for every observation) are well-defined.Hernán MA, Robins JM (2025). Causal Inference: What If. Chapman & Hall/CRC.
Kennedy EH (2019). Nonparametric causal effects based on incremental propensity score interventions. Journal of the American Statistical Association 114:645–656.
Díaz I, van der Laan MJ (2012). Population intervention causal effects based on stochastic interventions. Biometrics 68:541–549.
Díaz I, Williams N, Hoffman KL, Schenck EJ (2023). Non-parametric causal effects based on longitudinal modified treatment policies. Journal of the American Statistical Association 118:846–857.
---
title: "Intervention types in causatr"
code-fold: show
code-tools: true
vignette: >
%\VignetteIndexEntry{Intervention types in causatr}
%\VignetteEngine{quarto::html}
%\VignetteEncoding{UTF-8}
---
```{r}
#| include: false
knitr::opts_chunk$set(collapse = TRUE, comment = "#>")
```
causatr separates model fitting (`causat()`) from causal contrasts
(`contrast()`). The intervention --- the hypothetical change in treatment ---
is specified at contrast time, not fit time. This design lets you fit once and
contrast under many different interventions without re-estimating nuisance
models.
This vignette tours every intervention constructor, shows which estimators
support it, and demonstrates the calling pattern on real and simulated data.
## Intervention constructors at a glance
| Constructor | What it does | G-comp | IPW | AIPW | Matching |
|---|---|---|---|---|---|
| `static(v)` | Set treatment to `v` for everyone | All types | Binary, categorical | All types | Binary |
| `shift(d)` | Add `d` to observed treatment | Continuous | Continuous, count (integer `d`) | Continuous | --- |
| `scale_by(c)` | Multiply observed treatment by `c` | Continuous | Continuous, count (integer-preserving) | Continuous | --- |
| `threshold(lo, hi)` | Clamp treatment to `[lo, hi]` | Continuous | --- | --- | --- |
| `dynamic(rule)` | Apply a user-defined deterministic rule | All types | Binary, categorical | All types | --- |
| `ipsi(d)` | Multiply odds of treatment by `d` | --- | Binary | --- | --- |
| `stochastic(f)` | Draw treatment from user-supplied distribution | All types | --- | --- | --- |
| `grace_period(w)` | Delay treatment initiation by `w` periods (natural-history MTP) | Longitudinal, discrete | --- | --- | --- |
| `carry_forward()` | Carry the baseline treatment forward (LOCF; natural-history MTP) | Longitudinal, discrete | --- | --- | --- |
| `NULL` | Natural course (observed values) | All types | All types | All types | --- |
`grace_period()` and `carry_forward()` are **natural-history** modified
treatment policies (G-LMTPs): the intervened treatment at time $t$ depends on
the *counterfactual natural-value history* of treatment, so they cannot be
written as `dynamic()` rules (which see only the observed lag). They are
estimated by an augmented-data sequential regression and require a longitudinal
g-computation fit with a discrete treatment — see the
[longitudinal vignette](longitudinal.html) for runnable examples.
## Data: NHEFS
All examples use the **NHEFS** dataset (Hernán & Robins, 2025):
- **Binary treatment ($A$):** `qsmk` — quit smoking (0/1). Used for
`static()`, `dynamic()`, and `ipsi()` examples.
- **Continuous treatment ($A$):** `smokeintensity` — cigarettes smoked per
day. Used for `shift()`, `scale_by()`, and `threshold()` examples.
- **Outcome ($Y$):** `wt82_71` — weight change in kg.
- **Confounders ($L$):** sex, age, race, education, smoking-related
variables, exercise, activity, and baseline weight — common causes of
both $A$ and $Y$.
```{r}
#| message: false
library(causatr)
library(tinyplot)
data("nhefs")
nhefs_complete <- nhefs[!is.na(nhefs$wt82_71) & !is.na(nhefs$education), ]
```
## `static()` --- set treatment to a fixed value
The simplest and most common intervention. `static(1)` means "everyone is
treated"; `static(0)` means "no one is treated". For categorical treatments,
pass the level name as a string (e.g. `static("B")`).
### G-computation
```{r}
fit_gc <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "qsmk",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeintensity + I(smokeintensity^2) + smokeyrs + I(smokeyrs^2) +
factor(exercise) + factor(active) + wt71 + I(wt71^2),
model_fn = stats::glm
)
contrast(
fit_gc,
interventions = list(quit = static(1), continue = static(0)),
reference = "continue",
type = "difference"
)
```
### IPW
```{r}
fit_ipw <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "qsmk",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeintensity + I(smokeintensity^2) + smokeyrs + I(smokeyrs^2) +
factor(exercise) + factor(active) + wt71 + I(wt71^2),
estimator = "ipw",
model_fn = stats::glm,
propensity_model_fn = stats::glm
)
contrast(
fit_ipw,
interventions = list(quit = static(1), continue = static(0)),
reference = "continue",
type = "difference"
)
```
### AIPW (doubly robust)
```{r}
fit_aipw <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "qsmk",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeintensity + I(smokeintensity^2) + smokeyrs + I(smokeyrs^2) +
factor(exercise) + factor(active) + wt71 + I(wt71^2),
estimator = "aipw",
model_fn = stats::glm,
propensity_model_fn = stats::glm
)
contrast(
fit_aipw,
interventions = list(quit = static(1), continue = static(0)),
reference = "continue",
type = "difference"
)
```
### Matching
```{r}
fit_match <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "qsmk",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeintensity + I(smokeintensity^2) + smokeyrs + I(smokeyrs^2) +
factor(exercise) + factor(active) + wt71 + I(wt71^2),
estimator = "matching",
estimand = "ATT"
)
contrast(
fit_match,
interventions = list(quit = static(1), continue = static(0)),
reference = "continue",
type = "difference"
)
```
All four methods should give broadly similar estimates (methodological
triangulation). Discrepancies signal model misspecification in one or more
approaches.
## `shift()` --- add a constant to observed treatment
A modified treatment policy (MTP) that shifts each individual's observed
treatment by a fixed amount. For continuous treatments, this answers "what
would happen if everyone's exposure increased (or decreased) by `delta`
units?"
### G-computation
```{r}
fit_gc_cont <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "smokeintensity",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeyrs + I(smokeyrs^2) + factor(exercise) + factor(active) +
wt71 + I(wt71^2),
model_fn = stats::glm
)
res_shift_gc <- contrast(
fit_gc_cont,
interventions = list(reduce10 = shift(-10), observed = NULL),
reference = "observed",
type = "difference"
)
res_shift_gc
```
### IPW
Under IPW, `shift(delta)` uses a density-ratio weight: the ratio of the
treatment density evaluated at the inverse-shifted value to the density at
the observed value. The pushforward sign matters --- `shift(delta)` evaluates
at `A - delta`, not `A + delta` --- because the weight uses the inverse map.
```{r}
fit_ipw_cont <- causat(
nhefs_complete,
outcome = "wt82_71",
treatment = "smokeintensity",
confounders = ~ sex + age + I(age^2) + race + factor(education) +
smokeyrs + I(smokeyrs^2) + factor(exercise) + factor(active) +
wt71 + I(wt71^2),
estimator = "ipw",
model_fn = stats::glm,
propensity_model_fn = stats::glm
)
res_shift_ipw <- contrast(
fit_ipw_cont,
interventions = list(reduce10 = shift(-10), observed = NULL),
reference = "observed",
type = "difference"
)
res_shift_ipw
```
### Comparing shifts across methods
```{r}
#| fig-width: 7
#| fig-height: 3
#| fig-alt: "Point estimates and confidence intervals for a shift(-10) intervention under g-computation and IPW."
est <- data.frame(
method = c("G-comp", "IPW"),
estimate = c(
res_shift_gc$contrasts$estimate[1],
res_shift_ipw$contrasts$estimate[1]
),
ci_lower = c(
res_shift_gc$contrasts$ci_lower[1],
res_shift_ipw$contrasts$ci_lower[1]
),
ci_upper = c(
res_shift_gc$contrasts$ci_upper[1],
res_shift_ipw$contrasts$ci_upper[1]
)
)
tinyplot(
estimate ~ method,
data = est,
type = "pointrange",
ymin = est$ci_lower,
ymax = est$ci_upper,
xlab = "Method",
ylab = "Difference in weight change (kg)",
main = "shift(-10): reduce smoking by 10 cigs/day"
)
abline(h = 0, lty = 2, col = "grey40")
```
::: {.callout-note}
`threshold()` is **not supported** under IPW because the pushforward of a
continuous density under a boundary clamp is a mixed measure (point masses at
the bounds plus continuous density in the interior). Use
`estimator = "gcomp"` for threshold interventions.
:::
### Count treatment (Poisson / negative binomial)
For integer-valued count treatments, pass `propensity_family = "poisson"` or
`"negbin"` to use a count density model instead of the default Gaussian.
**DGM.** A Poisson-distributed count treatment (e.g. number of clinic
visits) confounded by $L$, with a linear outcome model.
$$
\begin{aligned}
L &\sim N(0,1) \\
A &\sim \text{Poisson}\!\bigl(\exp(0.5\,L)\bigr) \\
Y &= 2 + 1.5\,A + L + \varepsilon, \quad \varepsilon \sim N(0,1)
\end{aligned}
$$
True contrast: $\mathbb{E}[Y(A+1)] - \mathbb{E}[Y(A)] = 1.5$.
```{r}
set.seed(6)
n <- 3000
L <- rnorm(n)
A <- rpois(n, exp(0.5 * L))
Y <- 2 + 1.5 * A + L + rnorm(n)
count_df <- data.frame(Y = Y, A = A, L = L)
fit_pois <- causat(
count_df,
outcome = "Y",
treatment = "A",
confounders = ~ L,
estimator = "ipw",
propensity_family = "poisson",
model_fn = stats::glm,
propensity_model_fn = stats::glm
)
res_pois <- contrast(
fit_pois,
interventions = list(plus1 = shift(1), observed = NULL),
reference = "observed",
type = "difference"
)
res_pois
```
The shift delta must be integer --- `shift(0.5)` on a count treatment aborts
because `dpois()` returns 0 at non-integer arguments.
## `scale_by()` --- multiply observed treatment by a factor
A proportional MTP. `scale_by(0.5)` halves each individual's exposure;
`scale_by(2)` doubles it.
### G-computation
```{r}
res_scale_gc <- contrast(
fit_gc_cont,
interventions = list(halved = scale_by(0.5), observed = NULL),
reference = "observed",
type = "difference"
)
res_scale_gc
```
### IPW
Under IPW, `scale_by(c)` evaluates the density at `A / c` and includes the
Jacobian `|1/c|` in the density-ratio weight.
```{r}
res_scale_ipw <- contrast(
fit_ipw_cont,
interventions = list(halved = scale_by(0.5), observed = NULL),
reference = "observed",
type = "difference"
)
res_scale_ipw
```
## `threshold()` --- clamp treatment within bounds (g-comp only)
Caps treatment at a ceiling and/or floor. `threshold(0, 20)` means "nobody
smokes more than 20 cigarettes per day; those below 0 are set to 0".
```{r}
res_thresh <- contrast(
fit_gc_cont,
interventions = list(cap20 = threshold(0, 20), observed = NULL),
reference = "observed",
type = "difference"
)
res_thresh
```
This intervention is only available under g-computation. Under IPW, the
density-ratio weight is not well-defined at the boundary clamp points
(point-mass discontinuity in the pushforward density). causatr rejects
`threshold()` under `estimator = "ipw"` with an informative error pointing
at g-comp.
## `dynamic()` --- deterministic treatment rule
A dynamic intervention assigns treatment based on individual characteristics.
The `rule` function receives two arguments --- the full data and the observed
treatment vector --- and returns a vector of counterfactual treatment values.
### G-computation (binary treatment)
Assign quitting only to heavy smokers (> 20 cigarettes/day at baseline):
```{r}
res_dyn_gc <- contrast(
fit_gc,
interventions = list(
rule = dynamic(\(data, trt) ifelse(data$smokeintensity > 20, 1, 0)),
all_quit = static(1)
),
reference = "all_quit",
type = "difference"
)
res_dyn_gc
```
### IPW (binary treatment)
Under IPW, `dynamic()` on a binary treatment uses Horwitz--Thompson indicator
weights: `w_i = I(A_i = rule_i) / f(A_i | L_i)`. This is well-defined
because the treatment takes only two values and each individual's
counterfactual is deterministic.
```{r}
res_dyn_ipw <- contrast(
fit_ipw,
interventions = list(
rule = dynamic(\(data, trt) ifelse(data$smokeintensity > 20, 1, 0)),
all_quit = static(1)
),
reference = "all_quit",
type = "difference"
)
res_dyn_ipw
```
### G-computation (continuous treatment)
Dynamic rules on continuous treatments work under g-comp (predict-then-average
on the outcome model). Here, heavy smokers are reduced to 20 while others keep
their observed level:
```{r}
res_dyn_cont <- contrast(
fit_gc_cont,
interventions = list(
capped = dynamic(\(data, trt) pmin(trt, 20)),
observed = NULL
),
reference = "observed",
type = "difference"
)
res_dyn_cont
```
::: {.callout-note}
`dynamic()` on continuous treatment is **not supported** under IPW. A
deterministic rule on a continuous treatment is a Dirac per individual,
which has no Lebesgue density --- the density ratio is not defined. Rewrite
the rule as a smooth `shift()` or `scale_by()`, or use
`estimator = "gcomp"`.
:::
## `ipsi()` --- incremental propensity score intervention (IPW only)
Kennedy (2019)'s incremental propensity score intervention multiplies each
individual's *odds of treatment* by `delta`. It is a stochastic MTP that
never materialises a specific counterfactual treatment value --- instead,
it shifts the entire propensity score distribution.
The closed-form weight is:
$$w_i = \frac{\delta \cdot A_i + (1 - A_i)}{\delta \cdot p_i + (1 - p_i)}$$
where $p_i = P(A = 1 \mid L_i)$ is the fitted propensity.
IPSI is **binary-treatment, IPW-only** --- g-comp and matching cannot evaluate
it because there is no counterfactual treatment column to predict at.
### Doubling the odds of quitting
```{r}
res_ipsi_2 <- contrast(
fit_ipw,
interventions = list(double_odds = ipsi(2), natural = NULL),
reference = "natural",
type = "difference"
)
res_ipsi_2
```
### Halving the odds of quitting
```{r}
res_ipsi_half <- contrast(
fit_ipw,
interventions = list(half_odds = ipsi(0.5), natural = NULL),
reference = "natural",
type = "difference"
)
res_ipsi_half
```
### IPSI dose-response
Varying `delta` traces out a dose-response curve on the odds scale:
```{r}
#| fig-width: 7
#| fig-height: 4
#| fig-alt: "Dose-response curve showing the estimated effect of incrementally shifting the odds of quitting smoking."
deltas <- c(0.25, 0.5, 1, 2, 4)
ipsi_results <- lapply(deltas, function(d) {
res <- contrast(
fit_ipw,
interventions = list(shifted = ipsi(d), natural = NULL),
reference = "natural",
type = "difference"
)
data.frame(
delta = d,
estimate = res$contrasts$estimate[1],
ci_lower = res$contrasts$ci_lower[1],
ci_upper = res$contrasts$ci_upper[1]
)
})
ipsi_df <- do.call(rbind, ipsi_results)
tinyplot(
estimate ~ delta,
data = ipsi_df,
type = "pointrange",
ymin = ipsi_df$ci_lower,
ymax = ipsi_df$ci_upper,
xlab = expression(delta ~ "(odds multiplier)"),
ylab = "E[Y(IPSI)] - E[Y(natural)] (kg)",
main = "IPSI dose-response: effect on weight change"
)
abline(h = 0, lty = 2, col = "grey40")
```
At `delta = 1` the intervention is the natural course, so the contrast is
zero. Larger `delta` makes quitting more likely and the effect on weight
gain grows.
## `stochastic()` --- user-defined randomised intervention (g-comp only)
A stochastic intervention draws each individual's counterfactual treatment from
a user-supplied distribution that may depend on covariates. Unlike `dynamic()`,
which assigns a single deterministic value, `stochastic()` integrates over the
randomness by Monte Carlo: the g-formula is evaluated `n_mc` times, each with
an independent draw from the sampler, and the predictions are averaged.
The `sampler` function has the same signature as a `dynamic()` rule ---
`function(data, treatment)` --- but should return an independent random draw
each time it is called.
### Binary treatment: covariate-dependent randomisation
Assign quitting with probability that depends on smoking intensity:
```{r}
set.seed(42)
res_stoch <- contrast(
fit_gc,
interventions = list(
random_quit = stochastic(
\(data, trt) rbinom(nrow(data), 1,
plogis(-1 + 0.05 * data$smokeintensity)),
n_mc = 200L
),
all_quit = static(1)
),
reference = "all_quit",
type = "difference"
)
res_stoch
```
The estimate is a population average over both individuals *and* the stochastic
policy.
### Continuous treatment: random shift
Add a random perturbation (rather than a fixed shift) to each individual's
smoking intensity:
```{r}
set.seed(42)
res_stoch_cont <- contrast(
fit_gc_cont,
interventions = list(
noisy_reduce = stochastic(
\(data, trt) trt + rnorm(length(trt), mean = -5, sd = 2),
n_mc = 200L
),
observed = NULL
),
reference = "observed",
type = "difference"
)
res_stoch_cont
```
::: {.callout-tip}
**Choosing `n_mc`**: 100--500 is typical. The sandwich SE accounts for the MC
draws used at estimation time, so there is no need to inflate `n_mc` just for
inference. If point estimates visibly change when you re-run with a different
seed, increase `n_mc`.
:::
::: {.callout-note}
`stochastic()` is only supported under `estimator = "gcomp"` (point and
longitudinal). IPW and matching cannot evaluate it because the density ratio of
a user-defined stochastic policy is generally intractable.
:::
## `NULL` --- the natural course
`NULL` means "do not intervene; use the observed treatment values". It is the
natural reference for MTPs:
```{r}
res_multi <- contrast(
fit_gc_cont,
interventions = list(
reduce10 = shift(-10),
halved = scale_by(0.5),
cap20 = threshold(0, 20),
observed = NULL
),
reference = "observed",
type = "difference"
)
res_multi
```
## Combining interventions in one contrast
Both g-comp and IPW support multiple interventions in a single `contrast()`
call. All pairwise differences against the reference are computed, and the
variance-covariance matrix covers all interventions jointly.
### G-computation
```{r}
#| fig-width: 7
#| fig-height: 4
#| fig-alt: "Point estimates and confidence intervals for multiple continuous-treatment interventions under g-computation."
est <- res_multi$contrasts
tinyplot(
estimate ~ comparison,
data = est,
type = "pointrange",
ymin = est$ci_lower,
ymax = est$ci_upper,
xlab = "Comparison",
ylab = "Difference in weight change (kg)",
main = "Multiple interventions (g-comp)"
)
abline(h = 0, lty = 2, col = "grey40")
```
### IPW
```{r}
res_multi_ipw <- contrast(
fit_ipw_cont,
interventions = list(
reduce10 = shift(-10),
halved = scale_by(0.5),
observed = NULL
),
reference = "observed",
type = "difference"
)
res_multi_ipw
```
## Intervention support by estimator
The table below summarises which combinations are supported and which are
rejected with an informative error. The restrictions reflect genuine
architectural limits of the density-ratio framework, not implementation
gaps.
| Intervention | G-comp (point) | G-comp (longitudinal) | IPW | AIPW | Matching |
|---|---|---|---|---|---|
| `static()` | All treatment types | All treatment types | Binary, categorical | All types | Binary only |
| `shift()` | Continuous | Continuous | Continuous, count (integer delta) | Continuous | --- |
| `scale_by()` | Continuous | Continuous | Continuous, count (integer-preserving) | Continuous | --- |
| `threshold()` | Continuous | Continuous | --- (mixed pushforward) | --- | --- |
| `dynamic()` | All treatment types | All treatment types | Binary, categorical | All types | --- |
| `ipsi()` | --- | --- | Binary only | --- | --- |
| `stochastic()` | All treatment types | All treatment types | --- (intractable density ratio) | --- | --- |
| `NULL` | All | All | All | All | --- |
**Why some combinations are rejected:**
- **`threshold()` under IPW:** The pushforward of a continuous density under
a boundary clamp is a mixed measure --- point masses at `lo`/`hi` plus
continuous density in the interior. The density ratio is not well-defined.
- **`dynamic()` on continuous treatment under IPW:** A deterministic rule on
a continuous treatment is a Dirac per individual with no Lebesgue density.
- **`static(v)` on continuous treatment under IPW:** Nobody is observed
exactly at `v`, so the HT indicator `I(A = v)` is zero almost surely.
- **`ipsi()` under g-comp/matching:** IPSI shifts the propensity, not the
treatment value. There is no counterfactual treatment column to predict at.
- **`stochastic()` under IPW/matching:** The density ratio of an arbitrary
user-defined stochastic policy is generally intractable. Only g-computation
(Monte Carlo integration) supports it.
- **`static()` / `dynamic()` / `threshold()` / `ipsi()` on count treatments:**
Count treatments (enabled via `propensity_family = "poisson"` or `"negbin"`)
use `dpois()` / `dnbinom()` for the density ratio. `static()` is degenerate
(HT indicator is zero almost surely on a count distribution). `dynamic()` has
the same Dirac problem as on continuous. `threshold()` has the same mixed-
measure problem. `ipsi()` is Bernoulli-specific. Only `shift()` (integer
delta) and `scale_by()` (where `A / factor` is integer for every observation)
are well-defined.
## References
Hernán MA, Robins JM (2025). *Causal Inference: What If*. Chapman & Hall/CRC.
Kennedy EH (2019). Nonparametric causal effects based on incremental
propensity score interventions. *Journal of the American Statistical
Association* 114:645--656.
Díaz I, van der Laan MJ (2012). Population intervention causal effects
based on stochastic interventions. *Biometrics* 68:541--549.
Díaz I, Williams N, Hoffman KL, Schenck EJ (2023). Non-parametric causal
effects based on longitudinal modified treatment policies. *Journal of the
American Statistical Association* 118:846--857.